Public Channels
- # anonymous-question-box
- # ask-the-admin
- # day01-intro-genomics-biometric-model
- # day02-sem-study-design-phenomics
- # day03-quality-control-gwas
- # day04-heritability-and-gcta
- # day05-meta-analysis-prs
- # day06-genomicsem
- # day07-causal-modeling-mendelian-randomization
- # day08-sequencing-introduction-to-hail
- # day09-rare-saige
- # day10-pathway-and-gene-based-analyses
- # documentation
- # f-abcd-repronim-course-videos-instructions
- # f-creation-of-videos
- # general
- # python-intro
- # random
- # session-a
- # session-b
- # technical-support
Private Channels
Direct Messages
Group Direct Messages
@Jeff Lessem (he/him) has joined the channel
@matthew keller has joined the channel
@Katie Sheehan has joined the channel
@Jeff Lessem (he/him) - I noticed on the workshop website that it says “1-1.5 hours in live zoom breakout rooms…” I think some people will take this to mean that live sessions will last 1-1.5. I’d revise this to make it clear that we expect people to be in the live sessions for 2 hours per day, and this time will include live remarks from the instructor(s) and (mostly) working in small breakout rooms with other students.
Let me pin the link tot the list of tutors so ill find it in a few weeks: https://docs.google.com/spreadsheets/d/1W2nUsdp64zugXmUZ0roVazjeJ39OyASt0OvTpjBP2Dg/edit?usp=gmail#gid=0
@Lucía Colodro-Conde has joined the channel
hmmm… a total of 2 faculty have accepted the invite? That doesn’t bode well
and only one tutor has been added to the spreadsheet. I'll send out personal invites to the faculty next week.
That sounds like a good idea. Is it possible to give them a nudge at that time that the tutors also need to be on the spreadsheet?
I don't think it lets me add an extra message to an invitation.
@Kumar Veerapen has joined the channel
@Katrina Grasby has joined the channel
@Michael Neale has joined the channel
@Sarah Medland (she/her) has joined the channel
cant find the call link for in 10 min, where should I look for it?
katie sent it around. i’ll post it here in a sec
so the only two faculty NOT to complete the spreadsheet… <drum roll>… any guesses?
this should be the rowdy thursday facuty meeting at Jills restaurant right?
Yeash just brewed myself a decaf and am warpped in a blanket... different vibe...
Katie - can you make me an “owner” or whatever of the zoom call? I can’t create a poll
Matt- Both you and John are set as alternative hosts. I believe to make you the "owner" I will have to login to the meeting and set you as it. One second
okay you are now the host so you have the control 🙂
Can the various documents - workshop call, instructions for presenters etc be shared on here? Please?
*Thread Reply:* yes, sounds like a good idea. I think @Jeff Lessem (he/him) should create a (read only?) channel where we put all the official docs/correspondences
Hi Everyone,
The Next Boulder StatGen Workshop Call has been scheduled for Wednesday April 7th, at 3pm MTS. The instructions and link for the Zoom meeting are below.
Please let me know if you all have any questions, happy to help.
Thanks!
Katie Sheehan
Catlin Sheehan is inviting you to a scheduled Zoom meeting.
Topic: Boulder StatGen Workshop 4/7 @3PM MTS Time: Apr 7, 2021 03:00 PM Mountain Time (US and Canada)
Join Zoom Meeting https://cuboulder.zoom.us/j/96621272124
Meeting ID: 966 2127 2124 One tap mobile +12532158782,,96621272124# US (Tacoma) +13462487799,,96621272124# US (Houston)
Dial by your location +1 253 215 8782 US (Tacoma) +1 346 248 7799 US (Houston) +1 669 900 6833 US (San Jose) +1 301 715 8592 US (Washington DC) +1 312 626 6799 US (Chicago) +1 646 558 8656 US (New York) Meeting ID: 966 2127 2124 Find your local number: https://cuboulder.zoom.us/u/acDZzxSnbC
Join by SIP 96621272124@zoomcrc.com
Join by H.323 162.255.37.11 (US West) 162.255.36.11 (US East) 115.114.131.7 (India Mumbai) 115.114.115.7 (India Hyderabad) 213.19.144.110 (Amsterdam Netherlands) 213.244.140.110 (Germany) 103.122.166.55 (Australia Sydney) 103.122.167.55 (Australia Melbourne) 149.137.40.110 (Singapore) 64.211.144.160 (Brazil) 69.174.57.160 (Canada Toronto) 65.39.152.160 (Canada Vancouver) 207.226.132.110 (Japan Tokyo) 149.137.24.110 (Japan Osaka) Meeting ID: 966 2127 2124
We might (in total) have as many faculty + tutors as students this year! (Which is great in terms of being able to facilitate learning)
I'm at 66 faculty/tutors and around 160 students.
I wonder how this year’s workshop experience will shape how we conduct the workshop in future years…
The Workshop schedule is at https://www.colorado.edu/ibg/international-workshop/2021-international-statistical-genetics-workshop/workshop-2021-calendar-page
The Workshop syllabus is at https://www.colorado.edu/ibg/international-workshop/2021-syllabus
Hi, will the username and password be given later to connect to workshop.colorado.edu?. Thanks
*Thread Reply:* Yes, the username and passwords will be sent out soon.
*Thread Reply:* @Jeff Lessem (he/him) Thanks
Hey all the students are here! Hi I am Michel Nivard, I look forward to meeting you all during the practicals in perticular the GenomicSEM practical! If you have Q’s about the GenomicSEM videos come find me and the other faculty in the GenomicSEM slack channel!
Welcome students! We look forward to meeting you in a few weeks!
Hi everyone! Looking forward to ‘meet’ you - I’ll be hosting the Pathway sessions on the last day, feel free to reach out to me at any time here on slack though, as an alternative to the many coffee and tea chit-chat opportunities we would have had in Boulder. Have fun at the workshop!
Hi All! Looking forward to seeing you in a few weeks time. I'll be hosting the Causal modeling and Mendelian randomization sessions. Please feel free to post on slack/email me if you have any questions about the material!
Welcome to all students! I'll be hosting the Heritability sessions. Look forward to interacting with you soon.
Hi everyone! I look forward to meeting you all virtually soon 🙂 My research focuses on autism and other neurodevelopmental disorders. I'm relatively new to statistical genetics, and am working on a project involving polygenic risk scores, so am really excited to learn more about PRS, GWAS, and generally working with large genetic datasets!
Hi everyone! I'm Helen and my research focus is on using Mendelian randomization to investigate the possible causal effects of maternal environmental exposures during pregnancy on offspring health related outcomes. I'll be a tutor in the sessions on Quality Control and GWAS & Causal modeling and Mendelian randomization. Looking forward to meeting you!
Hi! I'm Jet, a PhD student based in the Karolinska Institute in Sweden, and my research focuses on eating disorders. Currently working on preparing the next GWAS for anorexia nervosa and binge eating together with my colleagues. Like Olivia, I'm relatively new to statistical genetics, and I'm looking forward to learn and meet you all!
Hi, I am Tong, and I am a postdoc at the dept of Medical Epidemiology and Biostatistics, Karolinska Institute. Several of my current projects are to investigate the comorbidities of asthma using population-based, twin, and post GWAS methods. I am relatively new to genetic epidemiology, and would like to learn more for example PRS, genomic SEM, and MR methods.
Hi! I am Giulio, PhD student from the University of Queensland, currently visiting at the University of Bristol (but based in London; long story). I have a background in behavioural genetics and I am doing my first PhD project on the relationship between birth weight and cardiovascular disease, using PRS with relatedness information. I am looking forward to meet everyone and I can't wait to learn more about statistical genetics! 😄
Hi everyone! I'm a PhD student in the psychology department at the university of Edinburgh, and I work on reading and language abilities. I'm particularly interested in GWAS, genomicSEM and using PRS to study GxE. I'm looking forward to getting started next week 🙂
Hi everyone, I am also a PhD student, interested in the genetics of brain structure and aging related processes. I am looking forward to a refresher on some of the gwas-based methods, but I am new to sequencing, causal modelling and gene-based analyses, which I am very excited about. Looking forward to meeting you next week!🔥
Hi everyone, I am postdoc at K.G.Jebsen Center for Genetic Epidemiology, Norwegian University of Science and Technology NTNU. I am currently working in exploring parental-offspring effect on several health outcomes. Looking forward to meet you all!
Hey everyone! I am a postdoc working at the Population Research Center at the UT Austin. I have a background in criminology and am currently studying the genetic etiology of externalizing behaviors. I can’t wait to meet you all and get started with the workshop!
Hi everyone! I'm a PhD student at the College of Medicine at Texas A&M University. I've worked on couple of projects so far about the genetic basis for neuroticism and the role of gene x environment interactions. I'm really new to statistical genetic, and I'm really looking forward to learning more and meeting you all!
Hello, I'm a PhD student at the University of Minnesota and I am in my last year! I primarily work with substance use (alcohol, tobacco, cannabis) and externalizing psychopathology phenotypes. My current project focuses on cannabis legalization in the United States. I am familiar with twin methodology but I am new to genomic methods, so I am hoping to learn some new skills there 🙂
Hi all! I'm a PhD student at McGill University in Montréal, Canada studying the genetics and genomics of amyotrophic lateral sclerosis. I'm looking at rare variants in patients, the interaction of polygenic risk with rare variants, repetitive genomic regions, and RNA expression changes in the nervous system. Really looking forward to this workshop and to broaden my knowledge of genetic methods!
Hi everyone! I'm a final year PhD student at Queen Mary University of London. I am currently using genomic SEM, GWAS & MR to study the shared genetics of dementia risk factors and how this influences dementia-related outcomes. I come from a non-genetic background (nursing and neuroscience) so am excited to attend this workshop to get a better grounding in the foundations of statistical genetics 🙂 really excited to meet you all!
Hi everyone! My name is Ravi and am a PhD Student at the University of Southern California. I'm working on genetics and neuroimaging of various diseases but mainly chronic pain. I'm really excited for this workshop and excited to meet everyone! 🧠🧬
Hi! I’m a PhD student at the University of Tokyo. My main interest is in methodological and study design aspects of genetic cohort studies such as participant selection, missing data, and bias resulting from those processes. I'm so excited to be able to learn the breadth of topics in modern statistical genetics!
Hello everyone! I am Grace and a Assistant Professor at the University of North Carolina at Chapel Hill School of Nursing. I'm new to statistical genetics, and hope to learn more about polygenic risk scores, GWAS data analysis, and gene expression analysis for my future project that involving individuals with eating disorders.
Hi everyone! I’m a PhD student at McGill University in Montreal, Canada studying the genetic architecture of autism and how rare and common variants interact to increase risk. I’m really looking forward to the workshop and learning how I can improve my current workflow and ensure statistical rigour in my analyses. Looking forward to meeting everyone, and feel free to connect with me on LinkedIn!
Hi, my name is Lydia. I’m an upcoming doctoral student at Colorado University Boulder. I’m currently working on a neuroimaging + placebo effects study. I have interest in genetic and neural contributors/markers of substance use phenotypes. I have some background in behavioral genetics and I look forward to learning more statistical genetic methods! Look forward to meeting everyone :)
Hi everyone! I am a lecturer at the University of Alicante. My research is focused on the areas of sleep and behaviour genetics. I am an active member of the Murcia Twin Registry. I have previously done some twin modelling and I very keen on learning more statistical genetic methods. Looking forward to meeting you all!
Hi everyone! I’m a first year PhD student at Edinburgh University, UK. I’m interested in the links between immune system function and major psychiatric disorders. I’m new to the field of statistical genetics. In my current project I’m using genomic SEM, PRSs and MTAG so very much looking forward to going over these in more detail and learning about other methods too.
Hi everyone! I'm a PhD student at the University of Iceland, but based in Sweden and therefore perform most of my work at Karolinska Institutet. My research focuses on psychological resilience in trauma exposed populations- and I'm particularly interested to learn more about GWAS, PRS and twin methodology, but excited also to learn about other methods. Looking forward to meeting you all!
Hello Everyone! I’m Mike Neale, I’m a professor at Virginia Commonwealth University and will be involved in the Day 2 session and as many others as I can. Looking forward to meeting everyone, sadly at a distance this year.
Hi everyone, I'm Barbara, a postdoc in neuroimaging genomics at the MPI for Psycholinguistics in Nijmegen, Netherlands. I'm working on projects related to human evolution but also currently involved in setting up a new ENIGMA study where this workshop comes in quite handy. Very excited for the next two weeks and to meet everyone!!
*Thread Reply:* Nice to see you here too!
Question for students & faculty: have you received my “Welcome to the workshop” email? We had some issues with it getting blocked going out so just making sure.
*Thread Reply:* great, thanks! So it seems like it went out to both faculty & students.
*Thread Reply:* I got it Matt! Thanks!
*Thread Reply:* I got it - it made it through our spam trap
*Thread Reply:* I actually didn't get this! It might be in a spam filter. when was it sent?
Hi everyone! I'm Luis, I will start my PhD at the University of Queensland in Australia in a couple of months. My research will focus on neurodegenerative or sleep disorders (I'm not sure yet). I'm interested in genomic SEM, GWAS and PRS. I am really excited to 'meet' you all soon! 😁
Hello all, I’m about to start a PhD on the genetics of bipolar and depression. I’m in Sarah Medland’s lab at the Queensland Institute of Medical Research, Australia. Looking forward to meeting you all, and a bit nervous about keeping up with the course - it is all very different from my previous work in public health!
Hi everyone! I am Giaco. I am a first-year Ph.D. student at the Max Planck School of Cognition. I am working on projects related to aesthetics, music, and, more broadly, hedonic value. I have worked and working with twin and genetic data. Needless to say, I am looking forward to getting to know people and methods in the next two weeks. 👋
Hi everyone! I am a PhD student in clinical psychology at Boston University. I work on a longitudinal study of Vietnam-era twins and my research interests include depression, suicidality, and cardiovascular health. I am excited to learn more about twin modeling and other statistical genetic methods!
Hello everyone! I'm Madhur. I'm about to start a Ph.D. in Quantitative Genetics w/ Mike Neale at Virginia Commonwealth University. I'm interested in the genetics, neuroimaging, and psychometrics of psychiatric disorders. Excited about learning new methods and getting to meet everyone. 👋
Hi all! I'll be a tutor in a couple of your sessions. I'm an Assistant Professor at Virginia Commonwealth University and am very excited to meet you. Please feel free to reach out if you are in the midst of prepping and have any questions related to R or others as you are getting ready (if I don't know the answer to your question, I generally know who does). No question too silly! ecprom@vcu.edu
hi everyone! my name is will. i am a postdoc at cambridge health alliance. i am excited about the workshop, and am mainly hoping to become a more critical consumer on genetics research on personality and better able to leverage genetic data to meet the identification assumptions of causal models. looking forward to meeting all of you!
*Thread Reply:* Random question - any chance your dad worked at National Technical Center/North Charles Research and Planning Group in Center Sq?
*Thread Reply:* yes he did work at north charles!
*Thread Reply:* did you work there as well?
*Thread Reply:* yes! from 1996 - 1998, right after my undergrad. I was his research assistant!
*Thread Reply:* haha no way! what a small world.
*Thread Reply:* sorry super slow reply - please tell him I said hello and send me best!
Hi everyone! My name is Jeremy Elman and I'm an Assistant Professor at UC San Diego. My research is focused on Alzheimer's and aging, primarily using neuroimaging and cognition. I have recently been incorporating genetic data so am excited to learn about new approaches that I may be able to apply. Looking forward to the next couple weeks (and already impressed with the onboarding we've received!)
*Thread Reply:* Great that you’re here, my friend!
*Thread Reply:* It's been a long time coming!
Hi everyone! I'm Guiomar, I'm finishing my PhD ar the University of Helsinki and I work in nutritional epidemiology and some genetic epìdemiology. I'm using data from the Finnish Twin Cohort FinnTwin16, I'm really excited to learn more about quantitative genetic methods and also I would like to know more about PRSs and Mendelian Randomization
*Thread Reply:* Several of us here work closely with Jaakko on a project looking at drinking in early midlife in FT16 - always nice to meet others working on other dimensions in these amazing cohorts!
Hello! I’m a PhD student at the Department of sociology and human geography at the University of Oslo. My research is on family sociological/demographic outcomes, i.e. marital status (marriage vs cohabitation) and fertility (age at first birth, total number of children). I’m particularly interested in polygenic risk scores and twin modeling. Looking forward to tomorrow!
*Thread Reply:* Hi Agnes! Nice to see a fellow researcher from UiO. I'm actually currently doing some work using Mendelian Randomization on fertility measurements (such at age at birth and total number of children) together with @Shannon D'Urso - hope to catch up during the next two weeks 🙂
*Thread Reply:* Hi Agnes! Part of my PhD involves looking at the relationship between fertility and endometrial cancer. It would be really interesting to have a chat with you about your work sometime 🙂
*Thread Reply:* Cool! Yes, that would be nice 🙂
Hi everyone! I am a postdoctoral fellow at PROMENTA Research Centre, Department of Psychology at the University of Oslo. My work focuses on risk factors that may make children and adolescents more susceptible to developing psychopathology. I am interested in polygenic risk scores and GCTA. Looking forward to e-meeting all of you!
*Thread Reply:* Hi Stella, Nice to see a fellow postdoc from UiO!
*Thread Reply:* Hi Gunn Helen, nice meeting you! I spotted a couple of other names from my department so I think there are many people from UiO. Let's see if we'll end up in the same breakout room for the practical session 🙂
Hi! I am Federico, I'm a first year PhD Student at the University of Murcia. My PhD has its focus on genetic factors involved in the covariation between sleep quality and depression, and because of that, I'm working along the Murcia Twin Registry. Furthermore, I've always liked statistical methods, so I am really looking forward to doing and enjoying this course!
Hi! My name is Laura and I’ve just started a PhD at the Nic Waals Institute. I’m working with data from the MoBa cohort and looking at within-family transmission of childhood mental health problems. I’m looking forward to the workshop!
Hi all! My name is Jacob and I’m a PhD student at the University of Minnesota’s Institute of Child Development. I’m interested in relationship influences (especially those happening in early life) on physical, cognitive, and mental health in adulthood. Some of my work uses data from the Minnesota Twin Registry and I’m looking forward to learning how best to leverage genetically informed studies to support/refute claims of causality!
*Thread Reply:* Fellow ICD alum here!
👋 Hi everyone! I am Geng Wang, a second-year PhD student of statistical genetics at the University of Queensland. My research uses genetics to understand the relationship between intrauterine environments and offspring cardiometabolic health. I am using different methods to improve the causal inference of developmental origins of cardiometabolic diseases, including Mendelian randomization, discordant twin study, and structural equation modelling. Looking forward to tomorrow. Twitter handle @JackGengWang
Hi all. My name is Andrew; I'm on clinical internship at Massachusetts General Hospital and will be starting as an Assistant Professor at CU Boulder this coming Fall. I'll be co-leading the Genomic SEM session next Monday with Michel Nivard. Really looking forward to meeting everyone tomorrow!
Hi all, my name is Shannon and I am a PhD student at the University of Queensland. My work focusses on using genetics to investigate causal relationships between the intrauterine environment and offspring health (in particular - neurodevelopment), as well as the more broad perinatal period and maternal health (eg endometrial cancer). Keen to get started 🙂
Hi everyone! Im Sarah Brislin- Im a postdoc at VCU working with Danielle Dick and the EDGE lab with an interest in genetic, neural, and environmental risk factors for externalizing behavior! Im excited to meet everyone and looking forward the the lectures
Hi everyone, My name is Isabella, I am a PhD student at the University of Copenhagen. I focus on the relation betweeen genetics and pharmaceutical treatment. Specifically between patients with autoimmune diseases that are treated with methotrexate. Look forward to the next two weeks, and meeting you all.
Hi everyone! My name is Yingjie and I’m a first-year PhD student at the Radboudumc and Donders Institute in the Netherlands. My current project involves using genetic profiling and neuroimaging techniques to study comorbidity and heterogeneity in psychiatric disorders. I’m looking forward to learning more about PRS, genomic SEM and sequencing. Looking forward to meeting you in a few hours!
Hi all, I am Perline, I am a PhD candidate at the Vrije Universiteit Amsterdam. I am interested in education, psychopathologies, noncognitive skills and how they relate to intergenerational transmission and inequalities. I am one of the tutor for the GenomicSEM session. Looking forward to it and have fun learning during the next weeks!
Hi everyone. I'm Lucía and I'm a first-year PhD student at the MPI for Psycholinguistics at the Language and Genetics Department. My research is focused on studying the common variation of social behaviour and related phenotypes. Looking forward to meeting you. 😃
Hello all! I am Kai and I am a PhD student at King's College London, studying the etiology of self-harm, particularly if there is etiological difference between suicidal and non-suicidal self-harm. Lovely to meet everyone here! 🙂
Hi everyone! I'm a PhD student at Uppsala University in Sweden. The focus of my research is on the etiology of different aspects of social behaviour and cognitive function in early infancy, using twin methodology. Looking forward to meeting all of you!
Hello everyone!! 🙂 My name Javier, I am a postdoctoral fellow at the University of Texas at Austin. My research focuses on the genetic architecture of aging-related phenotypes and cognition. I will be one of the tutors of the Genomic SEM session. Looking forward to meeting you!
Hello Everyone! I am Svetlana. I am a research assistant in the Psychiatric Genetics lab at the Queensland Institute of Medical Research. I am new to R, UNIX, and statistical genetics 🙂. I am excited about learning PRS and GWAS data analysis, and look forward to meeting you all!!
Hi I'm a research fellow from University of Tartu, Estonia :flag_ee:. I am interested in the causal effects of personality and cognition on obesity and health. As detailed GWASes for personality and cognition are missing, I am collecting personality and cognition in Estonian biobank. Once data is collected, I seek to use GWAS, Genomic SEM and Mendelian randomisation to understand links between behaviour and health.
Hi everyone! I am a PhD student at Florida State University, College of Criminology and Criminal Justice. The focus of my research is on the development of callous-unemotional traits in adolescents. Looking forward to meeting everyone!
Hi everyone! I’m a first year PhD student at VU University. My current project is to investigate whether adding eQTL and Hi-C data to gene- and gene-set analysis yields improved results. I’m really looking forward to learning more about Genomic SEM and pathway analyses.
Hello All! I'm Sage Hawn, a postdoctoral fellow in @ Boston University School of Medicine and the National Center for PTSD in Boston, MA, USA. I've done the Boulder workshop before but am looking forward to freshening up on various approaches after a year of clinical internship!
Hi everyone. I'm Marton Piroska, PhD student at Semmelweis University, Hungary. Our group mostly conducts twin research, so I'd like to broaden my knowledge of the statistical methods underlying these kinds of analyzes. Looking forward to meeting you all!
Hi everyone, I am Angelica Ronald, but go by Geli. I run the GEL lab at Birkbeck in London. I attended this course in 2002(!) and am here again as a refresher and to upskill in a few areas. Look forward to meeting everyone
Hi, I am Wikus and I am a postdoc at UCL, London. I'm currently investigating intergenerational risk of adhd, and am interested in modifiable risk factors of child and adolescent mental health problems.
Hi everyone! I'm Jessica and I'm a grad student at Universidade Federal de São Paulo in Brazil. My research focuses on psychiatric genetics and studying miRNA expression in adolescents with psychiatric disorders. I'm looking forward to meet you and to learn more about statistical genetics! =)
Hi everyone! I'm Maizy and I am a new grad student at CU Boulder. I am new to statistical genetics, but I am starting to develop some projects looking at the genetic architecture of anxiety disorders and other mental health problems. I am super excited to learn a lot, especially about Genomic SEM, and I can't wait to meet everyone!
Hi All! It's nice to see so many friends here (old and new) after 1.5 years apart! I am currently an Assistant Professor at Virginia Commonwealth University, and will soon be starting a new position at Rutgers University. I work on substance use disorders, with an emphasis on alcohol but also moving into cannabis. I am particularly interested in understanding the 'outside-the-skin' mechanisms through which genetic risk is transmitted in families, as well as social genetic effects (e.g., the influence of a spouse's genotype). I am also interested in using genetically-informative designs to strengthen inferences about how social-environmental factors shape substance use disorders (e.g., through the comparisons of siblings with different social-environmental exposures, or within-person designs). Although I wish we could be together in person, I really appreciate the thought and preparation that went into the format of this meeting
I am a postdoctoral fellow at Virginia Commonwealth Univ, working with Mike Neale. I will be helping to teach the course on day 2 (evening session). My background is in statistics and software development. I am one of the developers of OpenMx. Also, I am interested in the therapeutic use of psychedelic medicine.
Hi all. I’m a Bioinformaticien researcher at Qatar Genome program. I mainly do pipeline development and analysis for WGS. My background is in bioinformatics. Looking forward to learning about gwas and more statistical analysis. Also about QC.
Hi Everyone, I am Jinni Su, and I am an assistant professor of psychology at Arizona State University. I study gene-environment interplay and substance use among diverse populations, with an emphasis on how sociocultural factors, in conjunction with genetic factors, influence substance use outcomes among racial/ethnic minority youth. I look forward to learning more about genetics with you all in this workshop!
Hi everyone! I am a postdoc at the PROMENTA Research Centre, Department of Psychology at the University of Oslo. My work focuses on gene x environment interactions and parent-of-origin effects in developmental psychopathology. Looking forward to virtually meeting all of you!
*Thread Reply:* Hi Dinka! Great to meet fellow UiO researchers here 🙂 Hope to catch up during he next couple of weeks
*Thread Reply:* Hi Gunn-Helen, nice to meet you! 😊
Hi all, my name is Mischa and I am a PhD student at the University of Queensland. My work focusses on using genetics to investigate the relationships on chronic pain and osteoarthritis. Keen to get started and meet you all 🙂
Hi all, my name is Tong Chen and I am a PhD student in developmental psychology at Penn State University. My research interests are understanding genetic and environmental influences on internalizing problems and substance use in adolescence and early adulthood. Nice meeting you all!
Conor’s files can be found at the following location in the workshop cluster /faculty/conor/Boulder2021 If you need additional instructions: 1) copy files into your personal folder using your ssh client, 2) go to Rstudio and open/work from the folder where your newly copied data/code are located. You can use the following code in ssh client to copy everything into your folder: mkdir conor cd conor cp /faculty/conor/Boulder2021/** ./
Hi all, my name is Margherita and I am an Assistant Professor at Queen Mary university of London. My research focuses on investigating individual differences in cognitive development and education and I am particularly interested in the interplay between genes, environments and development. I will be tutoring on the GWAS QC and genomic SEM sessions. I look forward to seeing everyone over the next two weeks.
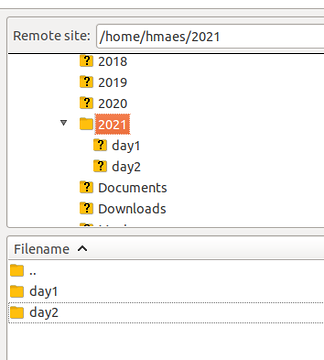
Hello! Files for Day 2 can be found at the following location. All subfolders should be copied. /faculty/hmaes/2021/day2/
If you need additional instructions. There are two ways you can copy files to your personal folder: METHOD 1 1) copy files into your personal folder using your ssh client. You can use the following code in ssh client to copy everything into your folder: mkdir day2 cd day2 cp -r /faculty/hmaes/2021/day2/** ./ 2) go to Rstudio on the server and open/work from the folder where your newly copied data/code are located. METHOD 2 1) Open up your SSH client. Then copy/paste, and run the line below. (shell script written by Tim Poterba [thank you Tim!] and edited for today). Then open your RStudio on the server and begin work. sh ~elizabeth/day2_prac.sh
*Thread Reply:* Thanks Elizabeth! As for method 1: I think you need to add -r to copy the directory contents to your folder. So:
cp -r /faculty/hmaes/2021/day2/** ./
*Thread Reply:* You're right. Thanks Jet! Will edit.
*Thread Reply:* @Elizabeth Prom-Wormley what will the code be for day 3 please (e.g., METHOD 1, above)? sorry, I wasn’t sure who to ‘@’ !!
*Thread Reply:* Hi @Laura the Day 3 code should be loaded soon. We're just trying to complete Session B. Once Session B is over, we'll get Day3 info up.
*Thread Reply:* Ah ok, thank you - sorry I’m jumping ahead!
*Thread Reply:* No worries. I love that you're super excited!
Hi everyone! I recently completed my PhD at the University of Gothenburg, Sweden, and will soon start a postdoc position at Karolinska Institutet in Stockholm, Sweden, with Cynthia Bulik, doing twin studies and GWAS on eating disorders, and the newly defined avoidant restrictive food intake disorder (ARFID) in specific. Looking forward to the course and to meeting everyone!
Hi everyone! I am a child psychiatry fellow at Yale and I am currently working on WES parent-child trio studies in a variety of childhood onset psychiatric conditions. Looking forward to continuing to meet everyone over the next 2 weeks and gaining some new skills in statistical genetics!
Once you enter your room today, please fill out the following survey. https://forms.gle/xUd8ZUocVxrF2DzTA
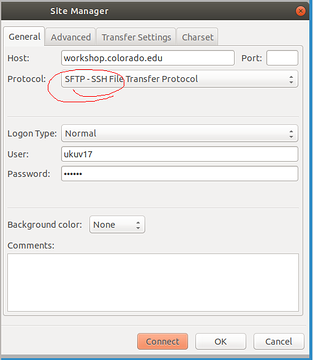
Who wants to have point-click interface for downloading files, use FileZilla for PC/Linux. Set it up as SFTP with SSH credentials. Then login, navigate one folder up to "home" and there you see the faculty members' folders
*Thread Reply:* Hi @Uku Vainik Could you direct my to some more detailed instructions about how to use FileZilla to transfer the course context to my own PC. Many thanks.
*Thread Reply:* hi try this one, ignore the one.com reference https://help.one.com/hc/en-us/articles/115005585709-How-do-I-connect-to-an-SFTP-server-with-FileZilla-
Hi Day 2 Session B! Files for Day 2 can be found at the following location. All subfolders should be copied. /faculty/hmaes/2021/day2/
If you need additional instructions. There are two ways you can copy files to your personal folder: METHOD 1 1) copy files into your personal folder using your ssh client. You can use the following code in ssh client to copy everything into your folder: mkdir day2 cd day2 cp -r /faculty/hmaes/2021/day2/** ./
2) go to Rstudio on the server and open/work from the folder where your newly copied data/code are located.
METHOD 2 1) Open up your SSH client. Then copy/paste, and run the line below. (shell script written by Tim Poterba [thank you Tim!] and edited for today). Then open your RStudio on the server and begin work. sh ~elizabeth/day2_prac.sh
Also, please be prepared to share one question that came up for you from the videos you watched while prepping for this session. See you soon!
*Thread Reply:* look in the faculty-general channel
I don't see that channel names are renamed to "day2 ..."
*Thread Reply:* "day02-" it starts with
*Thread Reply:* It's not updated in my slack. Maybe I need to reload?
*Thread Reply:* <#C01V93XE0JJ|day02-sem-study-design-phenomics>
*Thread Reply:* oh, hm, it looks like i need to join the channels
Hi Session B! Once you enter your room, fill out this survey https://forms.gle/5owbJcn5gDbvM7yc6
Hi @everyone - Below are 5 ways to get help during the workshop. I’ll pin this message and update it if needed: 4 ways to get help in REAL TIME:
- During breakout rooms, if there is a question that applies to two or more people in the room, use the “Ask for Help” button and a tutor will come by your room to help asap.
- During the breakout rooms, if you need help on an individual issue (something that applies only to you, such as being lost when your breakout roommates are not, a technical issue, etc.) you have the option of coming back to main room and we’ll try to get you 1-on-1 with a tutor. Please use this sparingly as we have limited tutoring resources. The first and preferred resolution to this is to first ask your roommates for help.
- Stick around after the live session, where you can hang out in the main room or your old breakout room, and some faculty/tutors may stick around and be able to provide help.
- You can try to direct message a tutor/faculty member and find a time/place to meet (e.g., in the usual zoom classroom). 1 way to get ASYNCHRONOUS HELP:
- At any time, ask a question on Slack in the relevant topic channel (now starting with “day01", “day02-” etc to make them easier to find) OR in the anonymous/random/general channel
@here if any European/Asian/Australias students have specific questions today regarding the last few practicals I’ll be online occasionally during the day and you can send me a message via chat if you want to talk. I’ll try to answer as soon as I am available. Obviously your Q might be one that others have so I encourage you to post in in a public channel and tag me (@Michel Nivard ) and/or the practical specific faculty member to grab our attention.
Postdoctoral Opportunity at the Institute for Behavioral Genetics, University of Colorado, Boulder IBG is one of the world’s leading institutes for genetic research on behavior. Internationally renowned research projects include the Colorado Adoption Project, the Colorado Twin Study and Longitudinal Twin Study, the Colorado Learning Disabilities Research Center, the Colorado Drug Research Center, and the Adolescent Brain and Cognitive Development (ABCD) Study. IBG is home to a DNA repository (about 40K samples) for research on human behavior, as well as studying behaviorally and genetically defined lines of selected, recombinant inbred, transgenic, and knockout-gene mice. Current research areas include aging, alcohol, behavioral development, brain structure and function, cognitive abilities and executive functions, drug abuse, evolution, neurodegenerative disease, nicotinic receptors, personality, psychopathology, reading and learning disabilities, and synaptic plasticity. Post-doctoral training opportunities are available through an NIH-funded T32 training grant administered through the National Institute on Mental Health (NIMH). Several investigators may serve as possible mentors for an outstanding candidate who wishes to further their training in behavior genetics. Projects include large human genetics studies, molecular genetics, and mouse behavioral studies aimed at uncovering and understanding genes related to mental health disorders. Successful applicants will have the opportunity to supervise and interact with junior graduate and undergraduate students. More information about IBG and the training program is located at www.colorado.edu/ibg. The position is restricted to US citizens or Green Card holders because of the funding source, but applicants from diverse and underrepresented backgrounds are encouraged to apply. Interested applicants should contact the faculty mentor with whom they would like to train and submit their complete application through CU Jobs at https://jobs.colorado.edu/jobs/JobDetail/?jobId=30468 The appointment could begin on July 1, 2021.
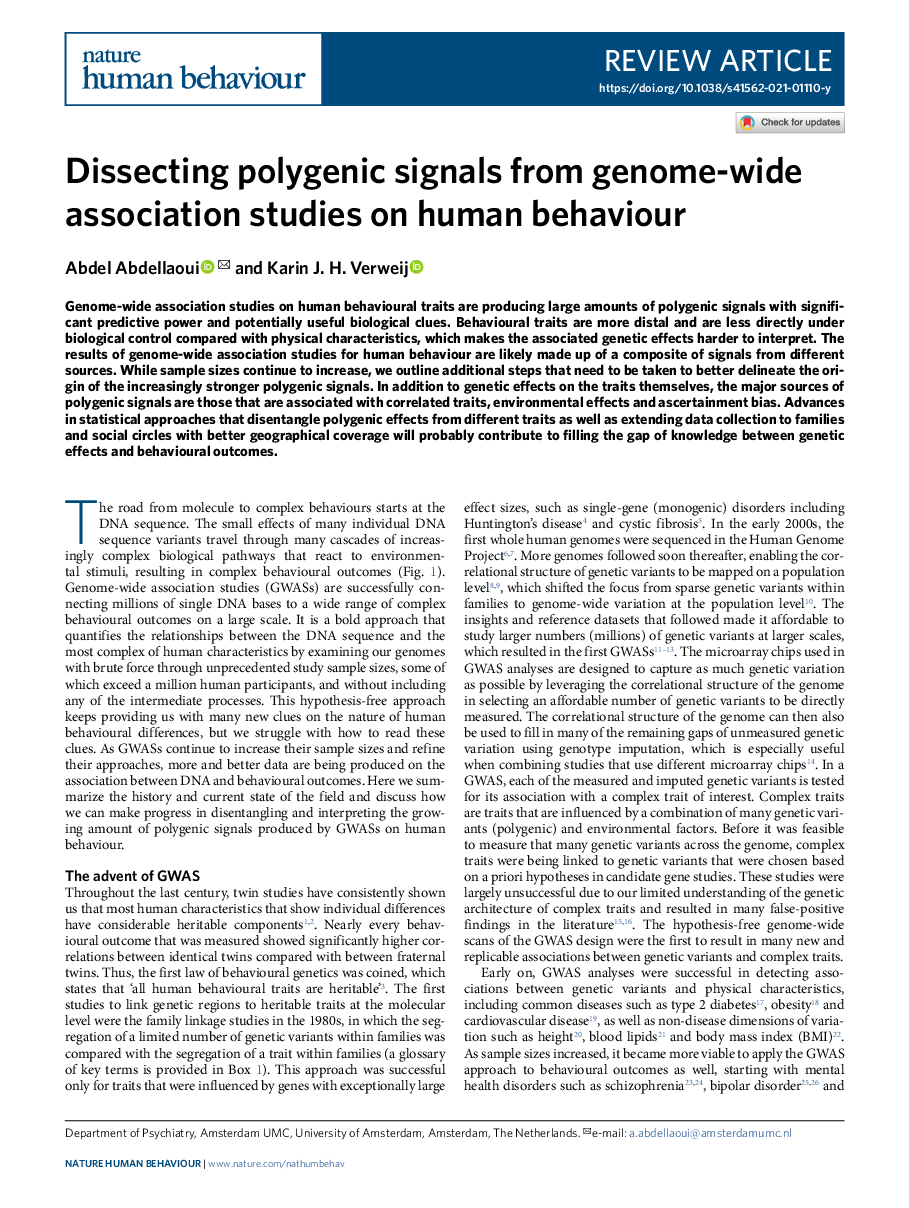
Hi all, I'm just going to use this channel to shamelessly (ok with a little shame) plug this recent review of ours on the history, current state, and future of GWASs of human behaviour. We wrote it to be accessible to a wide audience, such as the group of students of this course. Hope you enjoy it!
@channel Hi everyone. Notifications about talk/poster acceptance for the BGA conference were sent yesterday. Please check your spam folder if you didn't get an email.
Today's genetic trivia - the oldest rs number still in use is rs3 https://www.ncbi.nlm.nih.gov/snp/rs3#variant_details
We had one question in the tutorial today - how to you match your controls to your cases in your study? Anyone willing to share their best thoughts and tips on that?
*Thread Reply:* It’s a great question and was also asked anonymously — I took a stab at it here!
today's channel is <#C0204TTJ5LP|day04-heritability-and-gcta>
Hi all - we are hiring a research scientist in statistical genetics and psychiatric epidemiology. Further details in the ad attached here. Please let me know if you have any questions or would like to learn more!
Hi all - We are recruiting for a Post-Doctoral Associate position at IBG to work with Dr. Matthew Keller in collaboration with Dr. Peter Visscher. We are working on new approaches to understand the genetic and environmental architecture underlying complex traits. See ad here: https://jobs.colorado.edu/jobs/JobDetail/Post-Doctoral-Associate/30621
Do rare variants implicate the same genes as common ones? We’re doing a project on this and I was hoping for some crowd-source help. Is anyone aware of findings/papers that look at this question for trait X/Y/Z? E.g., a paper on autism showing that common SNPs are nearby genes already known to harbor highly penetrant rare variants? Thanks for your help all! But will poke @Benjamin Neale @Danielle
*Thread Reply:* Not exactly the same Q but a related one, do common and rare variants implicate the same geneest We chose this strategy because we knew power rwould be limited for individual genes: https://www.medrxiv.org/content/10.1101/2021.05.26.21257794v1
*Thread Reply:* Also: https://www.medrxiv.org/content/10.1101/2020.09.18.20192815v1 > We demonstrate that genes prioritized from common variant analyses of schizophrenia are enriched in rare variant risk, suggesting that common and rare genetic risk factors at least partially converge on the same underlying pathogenic biological processes.
Hi all! I plan on getting into our Zoom room 30 minutes before the session starts (A&B) if you think you need help and want to get a head start on copying any files or ensuring that you've got a good start to your practical. If you think need other additional help in advance, just send me a note. See you soon!
Here is a short video to show how to copy the files for day 5 (thanks Michel Nivard for the suggestion)
Maybe after a week of workshop, watching again the intro videos to R, unix, etc. help put things together ! 🤓
Good morning/evening/night later today (in 2.5 hrs) we are in the <#C020248GYLT|day06-genomicsem> channel, @Danielle will be in charge of assigning breakout rooms, directing tutors to students in need etc, I will present some slides (@Andrew Grotzinger will do so in the B session). Its going to be fun, we don't expect you to get it all done. we actually added assignments so you can get some further practice in after the workshop ends if you want to. I share this video which walks trough getting all the data for the session, your tutors for the A session are:
1 @Perline Demange 2 @Javier de la Fuente 3 @Jackson Thorp 4 @Andrea Allegrini 5 @Margherita Malanchini 6 @Mark Adams 7 @Gunn-Helen Moen
FYI: maybe you all already know (I saw some tutorials doing it), and I am stating the obvious, but for those who don’t, a nice hack in Rstudio to make life easier is to create named code subsections. You can do it just by following a commented line with four signs (i.e., ####). For me, it was game-changing. 😅
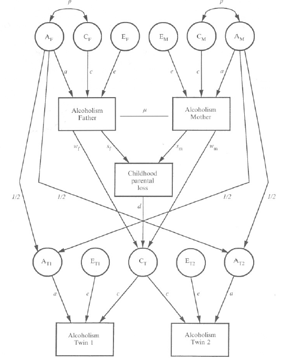
Might any twin modeling folks in the channel have a "twin family model with specified shared environment" script (or anything close to this that I could work from)? I am interested in doing something like this to understand the (measured) environmental mechanisms through which genetic risk is transmitted in families.
*Thread Reply:* I can, but there are some unresolved issues with this model. One is that we now know that cross-sectional mediation analyses yield biased results relative to the longitudinal data-generating mechanism used to simulate the data. Another problem here is that it’s not clear if parental loss occurs before or after the alcoholism (and selecting one or other flavor alone could distort things). To duplicate the model now, I’d probably start with Onyx, draw it and To Script it to get basic OpenMx code. In onyx I’d put in a two-headed arrow but in the mxPath() code for it I’d replace arrows=2 with arrows=0 (openMx syntax for a copath). I’d just duplicate the model for MZ twins, and move the path from AT2 to Alc Twin 2 to run from AT1 instead.
There are some gotchas still, in that there are non-linear constraints needed for i)residual variance for AT1 and AT2, ii) for the residual of CT, and the covariance between AT1 and CT. I think these were programed correctly for the article, and aren’t too hard to implement.
Hi @everyone
Just a heads up that the files used for tomorrow's practical session on Mendelian randomization can be found here: /faculty/davide/BOULDER2021/PRACTICAL1 /faculty/davide/BOULDER2021/PRACTICAL2 I would suggest creating a new folder in your own home directory called "MR" and then copying both directories to this new directory i.e.:
cd #Move to your home directory mkdir MR #Create a new directory called MR cd MR #Change to this directory cp -r /faculty/davide/BOULDER2021/PRACTICAL1 . #Copy PRACTICAL1 directory to the new MR directory cp -r /faculty/davide/BOULDER2021/PRACTICAL2 . #Copy PRACTICAL1 directory to the new MR directory
Look forward to seeing you tomorrow!
Dave
Please also be aware - if you've downloaded ahead of time. There is a typo in Practical2.R when setting the working directory it should be:
setwd("/home/****YOURNAMEHERE****/MR/PRACTICAL2/")
not PRACTICAL1. It's fixed now - so if you haven't downloaded it yet - don't worry about it!
For tomorrow, head over to this channel for Sequencing and Hail! 🙂 We posted an announcement to prep you for the excitement that is tomorrow’s workshop session @channel
@channel Behavior Genetics 2021 Annual Meeting Focus on applied methodology in Behavioral and Statistical Genetics Online 24 hour zoom meeting. Starts 7pm 28th June US-Eastern = midnight 28/29th June UK = 9am Aus-Eastern Live talks + interactive poster sessions. Join for the sections you are interested in (watch recorded sessions after) Registration by donation (suggested $24) For details see https://bga.org/current-meeting-2021/
MID COURSE SURVEY
Hi all. Thank you for taking the time to provide us feedback mid-course! This has been very helpful in shaping the faculty’s discussion this week of how future workshops might look, and has also been helpful in coming up with tweaks that might improve the little remaining time we have together. Also, a few of the specific ideas were really great (e.g., prac videos) and we’re strongly considering implementing them in future workshops.
Below, I summarize the feedback you provided. Please note that we will ask you again for a course evaluation at the end of the course, and it is really important to fill that one out because we report that feedback to the NIH and it helps determine whether this course can continue to be funded.
SUMMARY OF FEEDBACK: 61/71 (86%) satisfied or v. satisfied 60/72 (83%) watch most/all vids
Breakout room size preference (mean = 4.8). No strong pref for breakout rooms by skill (ppl with little experience wanted more experience in there).
Almost same breakdown of whether they would attend if virtual vs. in-person. However, in Q2.4 seems a bit more enthusiasm for in-person. Overall, most enthusiasm for a hybrid where some people attend in-person and others through video.
ASPECTS OF COURSE STUDENTS LIKE: • videos beforehand (better than live!) and great resource going forward • fantastic lectures; learning from leaders in the field • collective work • practicals • Slack Q&A • Medland’s prac approach • ability to fit virtual course into busy life with other demands • tutors helpful & accessible • low cost & no jetlag ASPECTS OF COURSE STUDENTS DISLIKE: • too much / too fast / not enough time / expects more background than I have • for some days, not enough effort to tie lectures to pracs [we’re working on this] • lack of networking • juggling time demands at home & in course • dealing with trivial syntax bugs [we’re working on this] • high variation in student experience • not enough time to discuss responsible conduct questions [we’re working on this] • disinterested zoom groups some days • no correction on closed captioning in some zoom lectures [we’re working on this] IDEAS: • having at least 1 experienced student per room • state learning goals at start of practical (what they will be doing & goals), then review/summarize what they did and what the point of it was [we’re working on this] • fewer topics, more depth • overall course flow diagram • tutors share screen more • videos of faculty members going through the prac step by step • have ppl figure out code rather than having code pre-written and running it • schedule time to watch vids with other students & fac • local hubs in hometowns
Hi Everyone. We will be taking a group photo in the day 9 practical session. We'll asking everyone to turn on their cameras so we can take a screen shot. Giving everyone advanced notice in case anyone wants to change their background. Thanks!
Hi Everyone, a sort of an open question for all but especially for faculty members. I'm looking for opinions/thoughts on the possibility of applying ML/AI approaches in the GWAS/genomics space. My transformational bioinformatics group in CSIRO from Australia is very interested in ML/AI so any suggestions/advice would be greatly appreciated. 🙂 Thanks!
*Thread Reply:* there is some discussion here: https://boulder-workshop.slack.com/archives/C01TLU7JAQK/p1623763267118400
*Thread Reply:* Thanks Tim! I really should have checked.
*Thread Reply:* Happy to help!
This may be a naïve question, but I was wondering if, in GWAS analyses, the alpha should always be set at p < 5e-8 -- which I believe is based on Bonferroni correction for approx. 10^6 independent tests. Or, should we set the alpha on a case-to-case basis, depending on the actual number of SNPs included - especially if the number of SNPs is orders of magnitude larger (or smaller) than 10^6?
*Thread Reply:* this isn’t naive and is an excellent question. For common variants, given the level of LD we observe in modern human populations, there are roughly 1 M independent SNPs across the genome and thus dividing .05 by 1M = 5e-8 is a justifiable alpha level. Of course, this is a rough approximation and differs a bit between populations (e.g., Africans have lower LD and thus need a slightly more conservative correction than Europeans, but this isn’t a huge difference and these thresholds, such as .05, are ultimately arbitrary anyway). When it comes to rare variants, the story is quite different. Rare variants tend to have much lower levels of LD, even with other rare variants, than do common variants with other common variants. Thus, as the MAF gets lower (e.g., if you’re using a v. large sequencing ref panel to impute your data down to, say, MAF = .0001, which is quite possible now), your GWAS will have more than 1M independent tests and you therefore need to use a more conservative correction. I’ll post a paper about this below. An attendant problem with a more conservative threshold for rare variants is lower power. As variants you want to test become increasingly rare, it may make sense to reduce teh number of tests performed by using gene-based or pathway-based association tests, or by using alternative approaches such as a Bayesian one that allows for weighting based on annotation.
Wu, Y., Zheng, Z., Visscher, P.M. et al. Quantifying the mapping precision of genome-wide association studies using whole-genome sequencing data. Genome Biol 18, 86 (2017). https://doi.org/10.1186/s13059-017-1216-0
Fadista J, Manning AK, Florez JC, Groop L. The (in)famous GWAS P-value threshold revisited and updated for low-frequency variants. Eur J Hum Genet. 2016;24:1202–5.

*Thread Reply:* to my mind, it depends on whether youd like find genome-wide significant SNPs or study-wide significant SNPs; if it is the first, then it does not matter how many SNPs youve tested, the genome-wide significance threshold will remain the same
*Thread Reply:* @Tetyana Zayats - perhaps I’m misunderstanding your point, but if you use 5e-8 in a GWAS on rare variants, you’ll get a lot of false positives, no?
*Thread Reply:* I`m sorry, I should have made it clear that my comment refers to common variants only.
*Thread Reply:* looking at the question, I`ve interpreted the GWAS as the studies on common variants only
*Thread Reply:* similarly, the 5E-08 is the genome-wide significance threshold for common variants only
*Thread Reply:* it was calculated using common variants only (if I`m not mistaken)
*Thread Reply:* Thank you both for the detailed explanations. @matthew keller, I was using the term "GWAS" rather loosely to include both common and rare variants. So, thanks for the relevant papers and methodological considerations. @Tetyana Zayats, thanks you for the distinctions between genome-wide and study-wide significance. That makes things clearer wrt. the impact (or not) of varying number of (common variant) SNPs across different studies.
*Thread Reply:* Great discussion here. I agree with Matt. Not a naive question at all. Just want to add this paper too as relevant. Note that this is based on sequenced data and not imputed SNPs as we normally do, which has an impact on false positive rates as shown in Wu et al. Still a relevant paper in particular to illustrate differences between ancestries. https://www.nature.com/articles/jhg201672
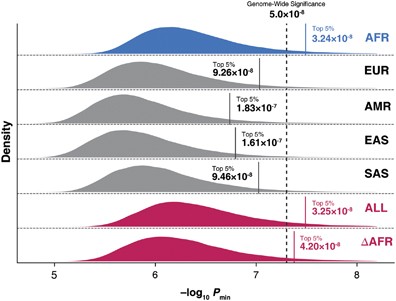
*Thread Reply:* Thanks so much @Loic Yengo for sharing this interesting paper!
How long will access and practice datasets be available after the workshop ends? Thx
*Thread Reply:* I can’t remember what we decided on this (or if we talked about it) but I’ll advocate that they be up indefinitely
*Thread Reply:* The cluster will be up at least through the weekend, but probably for another week or so. I'll be sending information on downloading your files sometime in the next few days.
@channel And that is officially a wrap! THANK YOU EVERYONE for being part of this years workshop!
Thank you to all of the faculty, tutors, hosts, and the incredible Jeff Lessem for all of their effort and help the past two weeks.
It was a great couple of weeks all! To old friends, it’s been good to catch up over the last two weeks, and to students, it’s been good to meet you to the degree possible. This workshop’s been a lot of mental effort and toil on the part of a lot of people. We weren’t sure what to expect, but in the end, I think it’s accomplished what we hoped (teach students) and more (begin the process of reconceptualizing what this workshop can be). Thank you again to all the fantastic faculty and tutors for your excellent work and the many many hours of your time. Thanks also to Jeff Lessem and the other staff here behind the scenes who made this workshop run seamlessly. And thanks to the excellent students - the level of engagement and questions this year was higher than at any in-person workshop I can remember. We hope that the online library of lectures and pracs we started this year will be a learning resource for you and your colleagues/mentees for years to come. Hope to see you at future conferences and workshops.
Hi Everyone There is still time to register for the online 2021 Behavior Genetics Meeting 28th/29th June if you are interested. You can register at: https://www.123signup.com/calendar?Org=BGA (registration is free - donation of $24 is suggested) You can find the program at: https://drive.google.com/file/d/1jcL4boL3Ts5wKhhUxPkQDnG4TgT-oL-T/view?usp=sharing best wishes Sarah